|
8/28/2023 0 Comments A human okazaki fragment This is also referred to as the "Trombone Model". 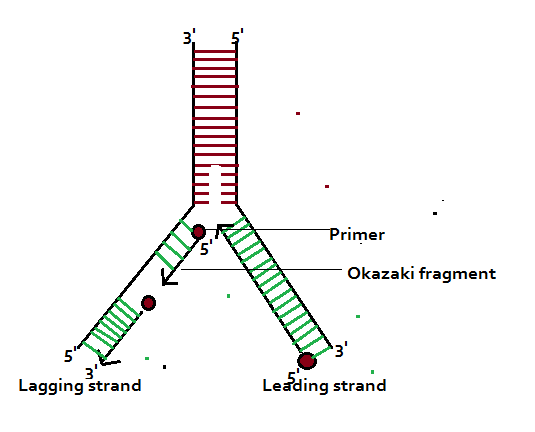
Correspondence to Accessibility StatementFor more information contact us noun, plural: Okazaki fragments 5-Ethynyl-2-deoxycytidine as a new agent for DNA labeling: detection of proliferating cells. Fast and accurate short read alignment with BurrowsWheeler transform. An enzyme called helicase unwinds the DNA by breaking the hydrogen bonds between the nitrogenous base pairs. Open Access articles citing this article. By submitting a comment you agree to abide by our Terms and Community Guidelines. Key points: DNA replication is semiconservative. Single-strand binding proteins coat the DNA around the replication fork to prevent rewinding of the DNA. You isolate a cell strain in which the joining together of Okazaki fragments is impaired and suspect that a mutation has occurred in an enzyme found at the replication fork. Binds to single-stranded DNA to avoid DNA rewinding back. FEN1-dependent cleavage of the short primer flap. Cutadapt removes adapter sequences from high-throughput sequencing reads. Genome-wide de novo L1 retrotransposition connects endonuclease activity with replication. Okazaki fragments are originally discovered by Reiji Okazaki, Tsuneko Okazaki, and their colleagues while studying replication of bacteriophage DNA in Escherichia coli in 1968. Peaks cloaked in the mist: The landscape of mammalian replication origins. When the replication fork reaches the end of the chromosome, however, there is (in many species, including humans) a short stretch of DNA that does not get covered by an Okazaki fragmentessentially, there's no way to get the fragment started because the. Okazaki fragment is a short newly-synthesized DNA fragment on the lagging template strand formed during DNA replication. Genome-wide identification and characterisation of human DNA replication origins by initiation site sequencing (ini-seq. Thus, Rad59 promotes fork progression when Okazaki fragment processing is compromised and counteracts PCNA-K107 mediated cell cycle arrest.Biochem. cdc9 rad59 double mutants did not alter PCNA ubiquitination but enhanced phosphorylation of the mediator of the replication checkpoint, Mrc1, indicative of increased replication fork stalling. To further understand how cells cope with nicks during replication, we utilized cdc9-1 in a genome-wide synthetic lethality screen and identified RAD59 as a strong negative interactor. Both enzymes reversed PCNA ubiquitination, arguing that the modification is likely triggered directly by nicks. To determine whether PCNA ubiquitination occurred in response to nicks or the lack of PCNA-DNA ligase interaction, we complemented cdc9 cells either with wild-type DNA ligase I or Chlorella virus ligase, the latter of which fails to interact with PCNA. In support of this notion, a pol30K107 mutation alleviated cell cycle arrest in cdc9 mutants. Most importantly, this signal is crucial to activate the S phase checkpoint, which promotes cell cycle arrest. The modification at K107 is catalyzed by the E2 variant Mms2 together with Ubc4 and the E3 ubiquitin ligase Rad5. cerevisiae as a model system, we uncovered a novel and conserved ubiquitination pathway that targets proliferating cell nuclear antigen (PCNA) at lysine 107 when DNA ligase I activity is inhibited. 
How cells monitor and suppress such accumulation of DNA damage that arises due to defective Okazaki fragment processing is unclear. An individual harboring DNA ligase I mutations exhibited growth retardation, sunlight sensitivity, severe immunosuppression and developed lymphoma, indicating a link between defects in Okazaki fragment maturation and cancer predisposition. 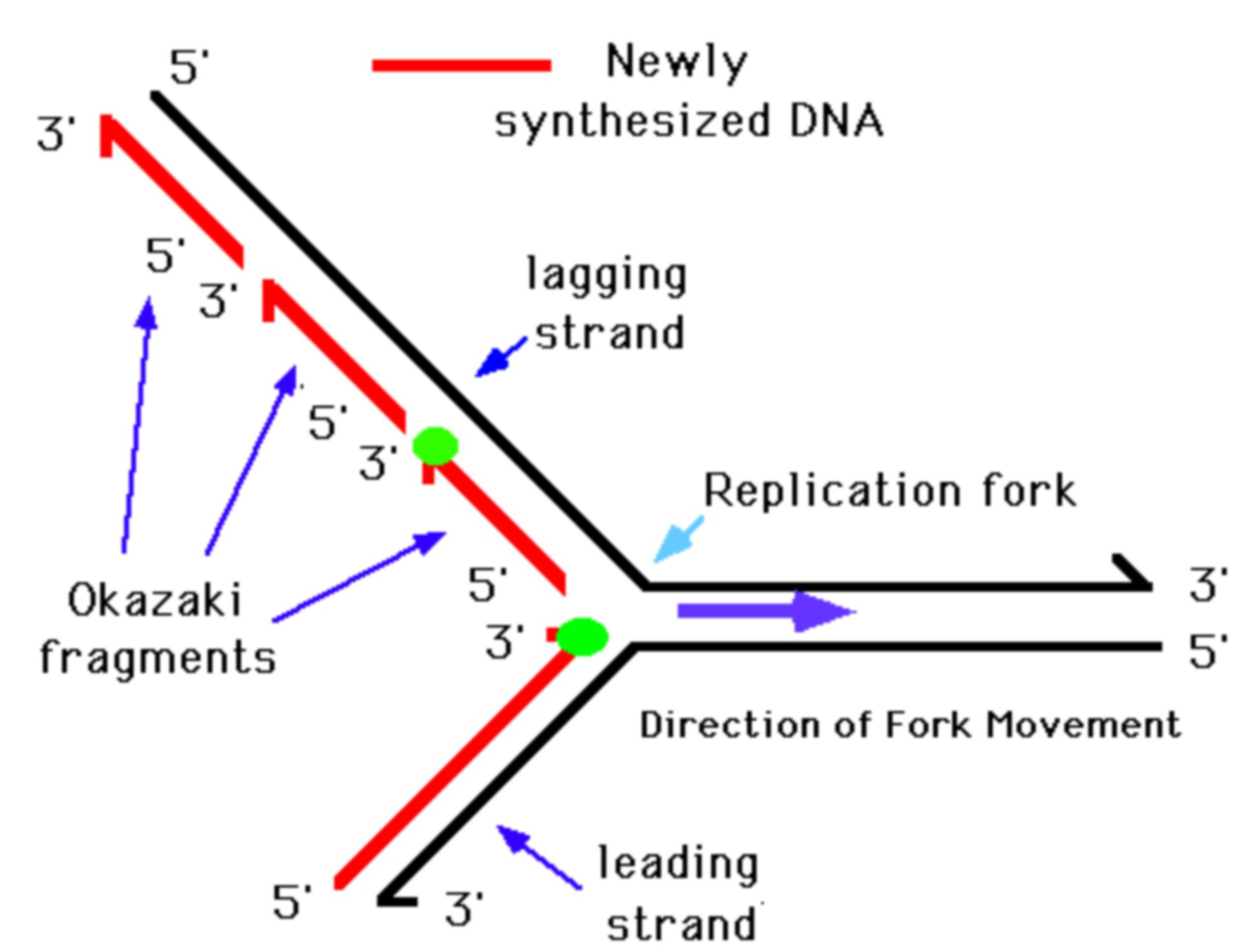
In humans, approximately 30 million Okazaki fragments are synthesized during every S phase and require further processing prior to DNA ligation. DNA ligase I, encoded by the CDC9 gene in Saccharomyces cerevisiae, is an essential enzyme that catalyzes the ligation of newly synthesized DNA on the lagging strand called Okazaki fragments. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
